News and Events> all events
- 02.-06.02.2020
SFB860-Retreat 2020, Marburger Haus, Hirschegg/Kleinwalsertal
- 04.-05.12.2019
SFB860-Symposium 2019, Max-Planck-Institut für biophysikalische Chemie, Göttingen
- 30.07.2019
Dr. Eike Christian Schulz, Abt. Dynamik in Atomarer Auflösung, Max-Planck-Institut für Struktur und Dynamik der Materie Hamburg
Press releases> Press release
- 12.06.2018
SFB an der Universität Göttingen verlängert. - 18.08.2014
Motoren-Umbau bei voller Fahrt. - 07.07.2014
SFB an der Universität Göttingen verlängert. - 03.06.2010
Universität Göttingen erhält neuen Sonderforschungsbereich.
Gallery
Contact
Spokesperson
Prof. Dr. Ralf Ficner 
Tel.: +49 (0)551-39-28618
Fax: +49 (0)551-39-28629
SFB860 Administration
Susanne van Beckum 
Tel.: +49 (0)551-39-28611
Fax: +49 (0)551-39-28629
Georg-August-Universität Göttingen
Abt. Molekulare Strukturbiologie
Justus-von-Liebig-Weg 11
37077 Göttingen
Introduction of SFB860
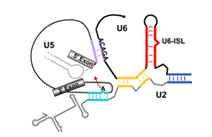
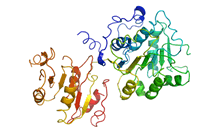
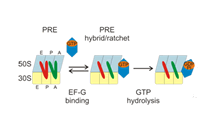
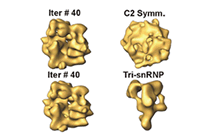
The dynamics of large macromolecular assemblies is a fundamental property that has great impact on their molecular and cellular function. This CRC will analyze the dynamics of several large macromolecular complexes with regard to their structures, their changes in conformation and composition, their interactions with other biological macromolecules, and with respect to their temporal and spatial location in the cell. Hence, the projects of the CRC focus on the determination of three-dimensional structures, the deduction of structure’s implication for the molecular and cellular function, the dynamic processes during assembly, remodelling and disassembly, the role of natively unfolded domains, the impact of posttranslational modifications on structure, function and dynamics, the conformational dynamics of functional complexes and their subunits, and kinetics and thermodynamics of macromolecular interactions. Studying the three-dimensional structure and the dynamics of single, average-size proteins is nowadays mostly a routine. However, due to the stunning complexity of large multi-protein- and ribonucleoprotein complexes, classical biochemical, biophysical and structure determination methods - when used exclusively - do not meet the requirements to comprehend such systems. Innovative strategies have to be developed for sample preparation, identification of macromolecular interaction networks, as well as for the analysis of compositional and structural dynamics. Hybrid approaches that combine single particle electron microscopy, crystallography, NMR spectroscopy, small angle X-ray scattering, single-molecule techniques and computational tools are required to generate testable atomic models of large macromolecular complexes. The CRC will bundle a wide spectrum of interdisciplinary competences present in Göttingen for the integrative analysis of a defined set of large macromolecular complexes, namely the spliceosome, the ribosome, the RNA degradosome, the mRNP locasome, the slam RNP, the mitochondrial translocase complex, nuclear export complexes, autophagy protein complexes, SLP-assembled signalosomes, the COP9 signalosome and the pyruvate dehydrogenase mulitenzyme complex. Since challenges and approaches are very similar for these functionally different macro¬molecular assemblies, the application of complementary biophysical and biochemical methods, as well as the exchange of experimental experiences and know-how will result in a significant added value for each project.
SFB 860 is funded by DFG (Deutsche Forschungsgemeinschaft)
1. funding period July 2010 - June 2014.
2. funding period July 2014 - June 2018.
3. funding period July 2018 - June 2022.
Publicationst; all Publications
- 04.2021
RNA helicase-mediated regulation of snoRNP dynamics on pre-ribosomes and rRNA 2'-O-methylation. - 04.2021
The RNA-binding protein FUS is chaperoned and imported into the nucleus by a network of import receptors. - 04.2021
The structure of Prp2 bound to RNA and ADP-BeF 3- reveals structural features important for RNA unwinding by DEAH-box ATPas. - 03.2021
A nuclear export sequence promotes CRM1-dependent targeting of the nucleoporin Nup214 to the nuclear pore complex. - 03.2021
Crystal structure of human CRM1, covalently modified by 2-mercaptoethanol on Cys528, in complex with RanGTP. - 01.2021
Ion type and valency differentially drive vimentin tetramers into intermediate filaments or higher order assemblies.